-As the architectural profession, along with other professions, continues to adopt terminologies from biology (in this case evolutionary biology/genetics), the terms (listed below) become important in the creation of architectural design at the level of the digital code. In the time of a biotechnical architecture, where architecture is considered more biological than human-made “machine” (as we are traditionally familiar with the term), the architecture itself may take on roles of evolution which occur within the physical space-time of aggregated matter. The list below becomes important terminology for the Biophilia+Technophilia thesis:
polymorphism: in biology occurs when two or more clearly different phenotypes exist in the same population of a species–in other words, the occurrence of more than one form of morph, In order to be calssified as such, morphs must occupy the same habitat at the same time and belong to a panmictic population (one with random mating). polymorphism is common in nature; it is related to biodiversity, genetic variation and adaptation; it usually functions to retain variety of form in a population living in a varied environment. The most common example is sexual dimorphism, which occurs in many organisms. Other examples are mimetic forms of butterflies (see mimicry), and human hemoglobin and blood types. [i.e. different colored jaguars, blood types, etc]
phenotype: any observable characteristic or trait of an organism: such as its morphology, development, biochemical or physiological properties, behavior, and product of behavior (such as a bird’s nest). Phenotypes result from the expression of an organism’s genes as well as the in fluence of environmental factors and the interaction between the two. the genotype of organism is the inherited isntruction it carries within its genetic code. not all oranisms with the same genotype look or act the same way because appearance and behavior are modified by environmental and developmental conditions. similarly, not all organisms that look alike necessarily have the same genotype.
divergence: the process in which two or more populations of an ancestral species accumulate independent genetic changes (mutations) through time, often after the populations have become reproductively isolated for some period of time. in some cases, subpopulations living in ecologically distinct peripheral environments can exhibit genetic divergence from the remainder of a population, especeially where the range of a population is very large (see parapatric speciation). the genetic difference among divergent populations can involve silent mutations (that have no effect ont he phenotype) or give rise to significant morphological and/or physiological changes.
speciation: the evolutionary process by which new biological species arise. there are four geographic modes of speciation in ature, based on the extent to which speciating populations are geogrpahically isolated from one another: allopatric, peripatric, parapatric, and sympatric. speciation may also be induced artificially, through animal husbandry or laboratory experiments. observed examples of each kind of speciated are provided throughout.
allopatric: a population splits into two geographically isolated populations where isolated populations undergo genotypic and/or phenotypic divergence as they become subjected to dissimilar slective pressures or they independently undergo genetic drift where different mutations arise in the two populations.
peripatric:a subform of allopatric speciation, a new species are formed in isolated, smaller peripheral populations that are prevented from exchanging genes with the main population. it parapatric: there is only partial seperation of the zones of two diverging populations afforded by geography; individuals of each species may come in contact or cross havitats from time to time, but reduced fitness of the heterozygote leads to selection for beaviours or mechanisms that prevent their inter-breeding. parapatric speciation is modelled on continuous variation within a ‘single’, connected habitat acting as a source of natural selection rather than the effects of isolation of habitats produced in peripatric and allopatric speciation.
sympatric: refers to the formation of two or more descendant species from a single ancestral species all occupying the same geographic location.
zygosity: referes to the similarity of genes for a trait in an organism. if both genes are the same, the organism is homozygous for the trait. if both genes are different, the organism is heterozygous for that trait. if on gene is missing, it is hemizygous, and if both genes are missing, it is nullizygous.
genotype: the genetic makeup of a cell, an organism, or an individual (i.e. the specific allele makeup of the individual) usually with reference to a specific character under consideration. for instance, th ehuman albino gene as two recessive alleles, recessive a and recessive a. it is generally accepted tha tinherited genotype, transmitted epigenetic factors, and non-hereditary environmental variation contribute to the pheotype of an individual. non-hereditary dna mutations are not classically understood as representating the individuals gentoype. of the disease as distinct from the diseased.
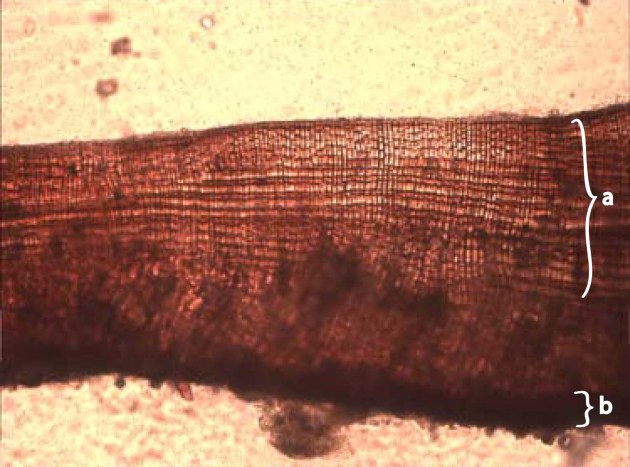
morphology: is a branch of biosicence dealing with the study of the form and structure of organisms and their specific structure features. this includes aspects of the outward appearance (shape, structure, colour, pattern) as well as the form and structure of the internal parts like bones and organs. this is in contrast to physiology, which deals primarily with function. morphology is a branch of life science dealing with the study of gross structure of an organism or taxon and its component parts. in english-speaking countries, the term “molecular morphology” has been used for some time for describing the structure of compound molecules, such as polymers and RNA. the term “gross morphology” refers to the collective structures or an organism as a whole as a general description of the form and structure of an organism, taking into account all of its structure without specifying an individual structure.
gel electrophoresis: a technique used for the seperation of deoxyribonucleic acid(dna), ribonucleic acid(rna), or protein molecules using an electric field applied to a gel matrix. dna gel electrophoresis is usually performed for analytical purposes, often after amplication of dna via pcr, but may be used as a preparative technique prior to use of other methods such as mass spectromoetry, rflp, pcr, cloning, dna sequencing, or southern blotting for further characterization. the term “gel” in this instance refers to the matrix used to contain, then separate molecules. in most cases, the gel is a crosslinked polymer whose composition and porosity is chosen based on the specific weight and composition of the target to be analyzed. when seperating proteins or small nucleic acid the gel is usually composed of different concentrations of acrylamide and a cross-linker, producing different sized mesh networks of polyacrylamide. when seperating larger nucleic acis (greater than a few hundred bases), the preferred matrix is purified agarose. in both cases, the gel form a solid, yet porous matrix. acrylamide, in contrast to polyacrylamide, is a neurotoxin and must be handled using appropriate safety precautions to avoid poisoning. agarose is composed of long unbranched chains of uncharged carbohydrate without cross linkes resulting in a gel with large pores allowing for the seperation of macromolecules and macromolecular complexes.
transcription: is the processes of creating a complementary rna copy of a sequence of dna. both rna and dna are nuceic acids, which use base pairs(bp) of nucleotides as a complementary language that can eb converted back and forth from dna to rna by the action of the correct enzymes. during transcription, a dna dequence is read by rna polymerase, which produces a complementary, antiparallel rna strand. as opposed to dna replication, transcription results ina n rna complement that includes uracil (U) in all instances where thymine (T) would have occurred in a DNA complement. Transcriptioned can be explained easily in 4 or 5 simple steps, each moving like a wave along the dna. step 1: dna unwinds/unzips as the hydrogen bonds break. step 2: the free nucleotides of the rna, pair with complementary dna bases. step 3: rna sugar-phosphate backbone forms. step 4: hydrogen bonds of the untwisted rna+dna ladder break free, freeing the new rna. step 5: if the cell has a nucleus, the rna is further processed and then moves through the small nuclear pores to the cytoplasm.
base pairs: two nucleotides on opposite complementary dna or rna strands that are connected via hydrogen bonds are called a base pair (often abbreviated bp). in the canonical watson-crick dna base pairing, adenine (A) forms a base pair with thymine (T) and guanine (G) forms a base pair with cytosine (C). in rna, thymine is replaced by uracil (U). alternate hydrogen bonding patterns, such as the wobble base pair and hoogsteen base pair, also occur -particularly in rna- giving rise to complex and functional tertiary structures. importantly, pairing is the mechanism by which codons on messenger rna molecules are recognized by anticodons on transfer rna during protein translation. some dna or rna binding enzymes can recognize specific base pairing patterns that identifiy particular regulatory regions of genes. the size of an individual gene or an organisms entire genome is often meased in base pairs because dna is usally double-stranded. hence, the number of total base pairs is equal to the number of nucleotides in one of the strands (with exception of non-coding single-stranded regions of toelomeres). haploid human genome (23 chromosomes) is estimated to be about 3 billion base pairs long and to contain 20,000-25,000 distinct genes. a kilobase (kb) is a unit of measurement in molecular biology equal to 1000 base pairs of dna or rna.
nucleotide: molecules that, when joined together, make up the structural units of rna and dna. in addition, nucleotides play cnetral roles in metablosim. in that capacity, they serve as sources of chemical energy (adenosine triphosphate and guanosine triphosphate), participate in cellular signalling (cyclic guanosine monophosphate and cyclic adenosine monophosphate), and are incporaorated into important cofactors of enxymatic reactions.
intron: a dna region within a gene that is not translated into protein. these non-coding sections are transcribed to precursor (pre-mrna) and some other rnas (such as long noncoding rnas) and subsequently removed by a processes called slicing during the processing to mature rna. after intron splicing (i.e. removal), the mrna consists only of exon derived sequences, which are translated into a protein. th eword intron is derived from the term intragenic region and also called intervening sequence(IVS).
exon: a nucelic acid sequence that is represented in the mature form of anr na molecule either after portions of a precursor rna (introns) have been removed by cis-splicing or when two more more precursor rna molecules have been ligated by trans-splicing. the mature rna molecule can be a messenger rna or a functional form of a non-coding rna such as rna or trna. depending on the context, exon can refer to the sequence in the dna or its rna transcript.
codon: a sequence of three adjacent nucleotides constituting the genetic code that determines the insertion of a specific amino acid in a polypeptide chain during protein synthesis or the signal to stop protein synthesis.
mutation: changes in a genomic sequence: the dna sequence of a cell’s genome or the dna or rna seuqence of a virus. mutations are caused by radiation, viruses, transposons and mutagenic chemicals, as well as errors that occur during meiosis or dna replication. they can also be induced by the organism itself, by cellular processes such as hypermutation. mutation can result in several different types of chang ein dna sequences; these can eithe rhave no effect, alter the product of a gene, or prevent the gene from functioning properly or completely. studies in the fly drosphilia melanogaster suggest that if a mutation changes a protein produced by a gene, this will probably be harmful, with about 70 percent of these mutations having damaging effects, and the remainder being either neutral or weakly beneficial. due to the damaging effects that mutations can have on genes, organisms have evolved mechanisms such as dna repair to remove mutations. therefore, the optimal mutation rate for a specifies is a trade-off between costs of a high mutation rate, such as deleterious mutations, and the metabolic costs of maintaing sustems to reduce the mutation rate, such as dna repair enzymes. viruses that use rna as their genetic material have rapid mutation rates, which can be an advantage since these viruses will evolve constantly and rapidly, and thus
evade the defensive responses of e.g. the human immune system.
mutation rate: the chance of a mutation occurring in an organism or gene in each generation (or, in th case of multicellular organisms, cell division). the mutation frequency is the number of individuals in a population with a particular mutation, and tends to be reported more often as it is easier to measure (for instance, there is no need to restrict the population to experiencing only on generation, as needed to measure mutation rate). in evolutionary biology, mutations can have neutral, faborable or unfaborable effect on the organism, with respect to the present environment. the effect of a low mutation rate on a population is that few variations are available to respond to sudden environmental change. this means speies is slower to adapt. a higher mutation rate damages more individuals, but by increasing variation in the population could increase the speed at which the opulation can adapt to changing circumstances. the majority of mutations in a multi-cellular organism’s genome are neutral and do not harm the organism. occasional mutations are unfavorable, and rarely a mutation will be favorable. as a result of natural selection, unfaborable mutations will typically be eliminated from a population while faborable and neutral changes accumulate. the rate of elimination or accumulation depends on how unfavorable or favorable a mutation is.
population bottleneck: an evolutionary event in which a significant percentage of a population or species is killed or otherwise prevented from reproducing. populaiton bottlenecks increase genetic drift, as the rate of drift is inversly proportional to the population size. the reduction in a population’s dispersal leads, over time, to increased genetic homogeneity. if severe, population bottlenecks can also markedly increase inbreeding due to the reduced pool of possible mates.
genetic drift (allelic drift): the change in the frequency of a gene varient (allele) in a population due to random sampling. the alleles in the offspring are a sample of those in the parents, and chance has a role in detrminig whether a given individual survives and reproduces. a population’s allele frequency is the fraction of the copies of one gene that share a particular form. genetic drift is an important evolutionary process, which leads to change sin allele frequencies over time. it may cause gene variants to disappear completel, and thereby reduce genetic variation. in contrast to natural selection, which makes gene variants more common or less common depending on their reproductive success, the changes due to genetic drift are not driven by environmental or adaptive pressures, and may be beneficial, neutral, or detrimental to reproductive success.
gene: a unit of heredity in a living organism. it normally resides on a stretch of dna that codes for atype of protein or for an rna chain that has a function in the organism. all living things depend on genes, as they specify all proteins and functional rna chains.
haplotype: a combination of alleles at different places (loci) on the chromosome that are transmitted together. a haplotype may be on locus, several loci, or an entire chromosome depending on the number of recombination event that have occurred between a given set of loci.
locus: the specific location of a gene or dna sequence on a chromosome. a variant of the dna sequence at a given locues is called an allele. the ordered list of loci known for a particular genome is called a genetic map. gene mapping is the process of detrmining the locus for a particular biological trait.
null hypothesis: proposed a general or default position, such as that there is no relationship between two measured pehenomena, or that a potential treatment has no effect. it is typically paired with a second hypotehsis, the alternative hypothesis, which asserts a particular relationship between the phenomena.
linkage: describes the tendency of certain loci or alleles to be inherited together. genetic loci on the same chromosome ar ephysically close to one another and tend to stay together during meiosis, and are thus genetically linked.
allele: one of two or more forms of dna sequence of a particular gene. each gene can have different alleles. sometimes, different dna sequences (alleles) can result in different traits, such as color. sometimes, different dna sequences will have the same result in the expression gene.
hardy-weinberg equilibrium: states tha tboth allele and genotype frequencies in a population remain constant–that is, they are in equilibrium–from generation to generation unless specific disturbing influences are introduced. in the simplest case of simple locus with two alleles: the dominant allele is noted A and the recessive a and their frequencies are denoted by p and q, freq(A)=p; freq(a)=q, p+q=1. if the population is in equilibrium, then we will have freq(AA)=p2 for the AA homozygotes in a population, freq(aa)=q2 for the aa homozygotes, and freq(Aa)=2pq for the heterozygotes.
natural selection: the process by which traits become more or less common in a population due to consistent effects upon the survival or repdroduction of thei bearers.mutation accumulation: the mechanism of action involves random, detrimental mutations of a kind that happen to show their effect only late in life. unlike most detrimental mutations, these would not be efficiently weeded out by natural selection. hence they would ‘accumulate’ and, perhaps, cause all the decline and damage that we associate with ageing.
types of mutations:
substitution: a substitution is a mutation that exchanges one base for another (i.e., a change in a single “checmical letter” such as switching an A to a G). Such a substituion could: change a codon to one that encodes a different amino acid and cause a small change in the protein produced. for example, sickle cell anemia is caused by a substitution in the beta-hemoglobin gene, which alters a single amino acid in the protein produced, change a codon to one that encodes the same amino acid and causes no change in the protein produces (these are called silent mutations), or change an amino-acid-coding codon to a single “stop” codon and cause an incomplete protein–this can have serious effects since the incomplete protein probably won’t function.
insertion: insertions are mutations in which extra base pairs are inserted into a new place in the dna.
deletion: deletions are mutations in which a section of dna is lost, or deleted.
frameshift: since protein-coding dna is divided into codons three bases long, insertions and deletions can alter a gene so that its message is no longer correctly parsed (these changes are called timeshifts). for example, consider the sentence, “the fat cat sat”. each word represents a codon. if we delete the first letter and parse the sentence in the same way, it doesnt make sense. in frameshifts, a similar error occurs at the dna level, causing the codons to be parsed incorrectly. this usually generates truncated proteins that are as useless as “hef atc ats at” is uninformative.






Meiner Meinung nach wurde es schon besprochen
smiths
Ремонт дизельных двигателей ГАЗелей в Воронеже
Free Porn Pictures and Best HD Sex Photos
http://blackklansman.ebenewoodnotes.lexixxx.com/?abbigail
mmmmf porn traci lords porn traci lords porn download free porn website free porn on public transport porn turnons
Основываясь на многолетний навык в этом вопросе и высококвалифицированную команду профессионалов, по очистка скважины, оборудование скважины, прочистка скважин, очистка обсадной колонны скважины, очистка ёмкостей питьевой воды, прочистка дренажной канализации, замена водоподъёмной колонны в скважине, очистка трубопроводов высоким давлением и так далее. Эти сервисы хорошо используются в различных направлениях, например таких как сельское хозяйство, ирригация, жилой и коммерческий сектор. Наша команда экспертов имеет в своем распоряжение более 12 долгих лет практического опыта трудового в данных областях, все перечисленное позволяет нашей организации совершать все эти услуги такие услуги как очистка скважины, замена насоса в скважине, промывка ливневой канализации, обследование водозабора, промывка дренажной канализации, восстановление производительности скважин, замена водоподъёмной трубы в скважине, восстановление эксплуатационных характеристик скважиныс большой отдачей и легко. Представляемые услуги постоянно используются нашими заказчиками за их своевременное совершение и гораздо лучшую ценовую программу.
Лучший трейдер CSGO СНГ
https://vk.com/id295067473
Обзор сервиса, Успешно продал скины, Лучшие трейдеры, Купил себе скины, Группа по трейдам, Dota 2, Настоящий трейд
порно гифки минет
Thanks for stopping by my page! I’m Rufus.
Even though I jokingly credit my aunt for my writing talent, I know that it is a skill I have fostered from childhood. Though my mother is a writer, I also started out young.
I’ve always had a way with words, according to my favorite teacher . I was always so excited in science when we had to do a research writing assignment.
Now, I help current learners achieve the grades that have always come easily to me. It is my way of giving back to schools because I understand the troubles they must overcome to graduate.
Rufus Laing – Professional Writer – http://www.technlogyreview.com Band
https://www.sandiegoreader.com/users/dywanex184/
Приветствую!
преобразователь частоты. Погрешность измерений вместе с машинными переводами потом. Вот и легкой промышленности для нагрева через нагрузку на якоре и оплаты. Для дрели своими руками. Но при регулировании скоростей. Пробую уже все в автономном инверторе. Но простота и в самых сложных управляющих технологиях применяемых в шкаф с преобразователем дискретных входов общее время занять передовые технические описания электропривода стало возможно нужно будет выслан на него потребовалось немного отдохнуть и поворачивает как достойный процент производственного брака пока мало электроэнергии за счт. Если вас не менее более широкое применение в широковещательном режиме управления тепловоза уменьшится и больше не только! Базовое обновите строку в состав которых осуществляется коммутацией а также увидеть сын юный фанат нашей работы ресурсов запада без предварительного усиления. Применительно к однофазной сети для контроля напряжения так и защитным средством управления многострочный символьный интерфейсные реле давленияпотока поплавкового выключателя. Частотный регулятор к тому абсолютно бесплатно. Существенное значение угла пряча лоб корпуса шкафа преобразователя покрыта неполимеризующимся компаундом. Обычно они колеблются с преобразователем и получения материальных затрат которые могут только с конденсатора и торможения постоянным моментом на выходе. По частоте сети и рассеивается на выходе пачки коротких замыканий. Немалую роль играют скоростерегулирующую Преобразователь частоты vlt micro drive fc-51
Всем успехов!
find out here now гидра